Fragment network basics

In my last post I described how the fragment network can be used as a key part of a virtual screening project by providing allowing your initial hits to be expanded out to a large number of candidates to screen. In this post we describe how the fragment network works and why is more ‘chemically intuitive’ that traditional fingerprint based similarity search.
For the fullest understanding I’d suggest you read the Astex paper. I try to summarise the key points here.
The approach involves breaking the input molecules into fragments by breaking bonds that attach functional groups to rings, or attach a ring to another ring. As an example consider 4-hydroxy biphenol:
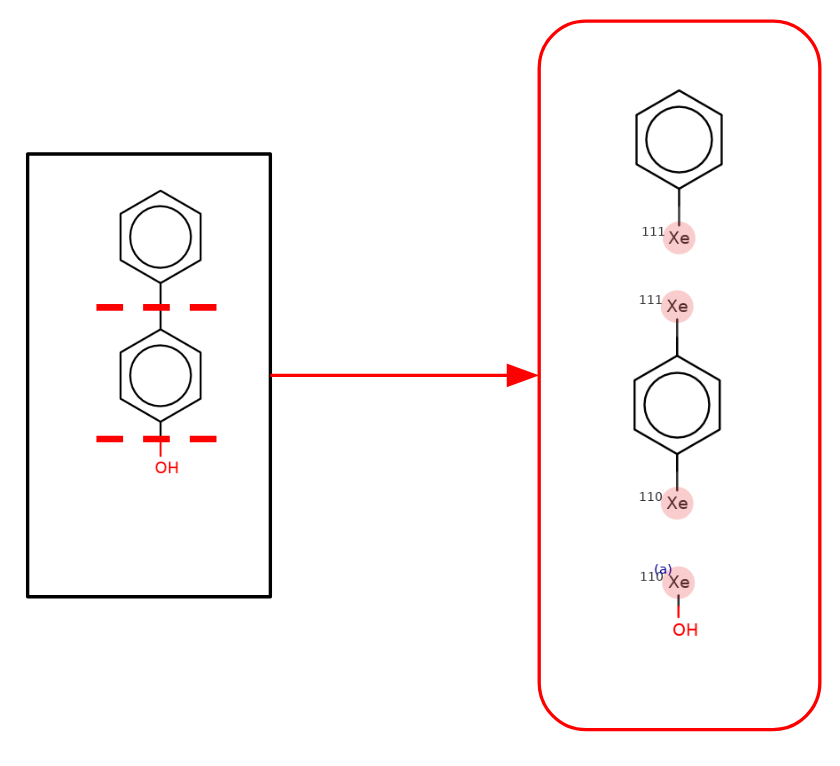
The molecule is split as shown resulting in 3 fragments. Xenon atoms are used to mark the sites of fragmentation. Then those 3 fragments are recombined in an “all but leave out one” manner to generate a number of child molecules. In addition in the special case of a ring-ring split (rings joined with a single bond or through a spiro atom) those two fragments are included as a child. That results in the following 4 child molecules:
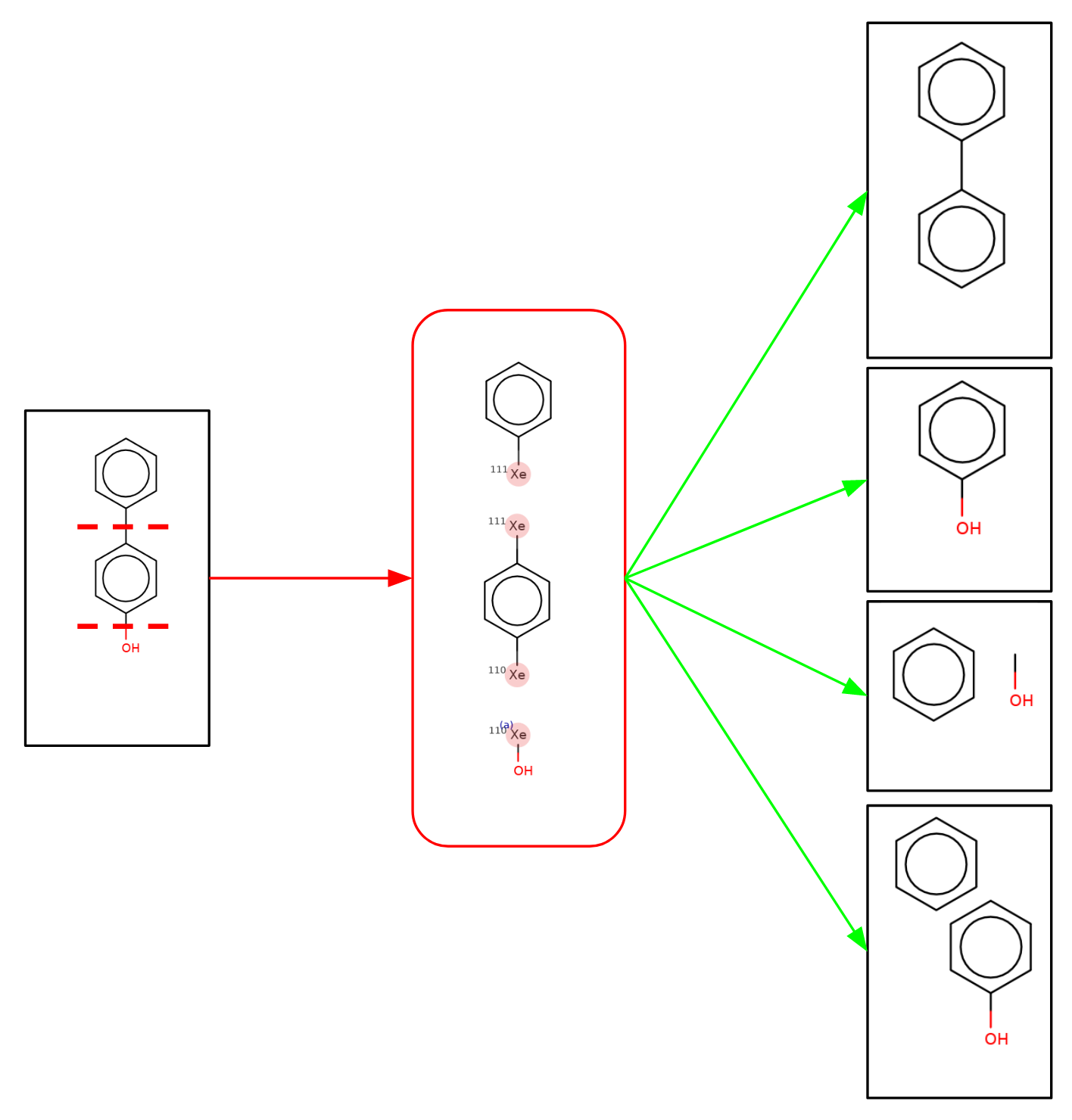
Those child molecules are in turn fragmented and re-combined until there are no more fragments. For our example this results in the following directed acyclic graph of parent and child molecules:
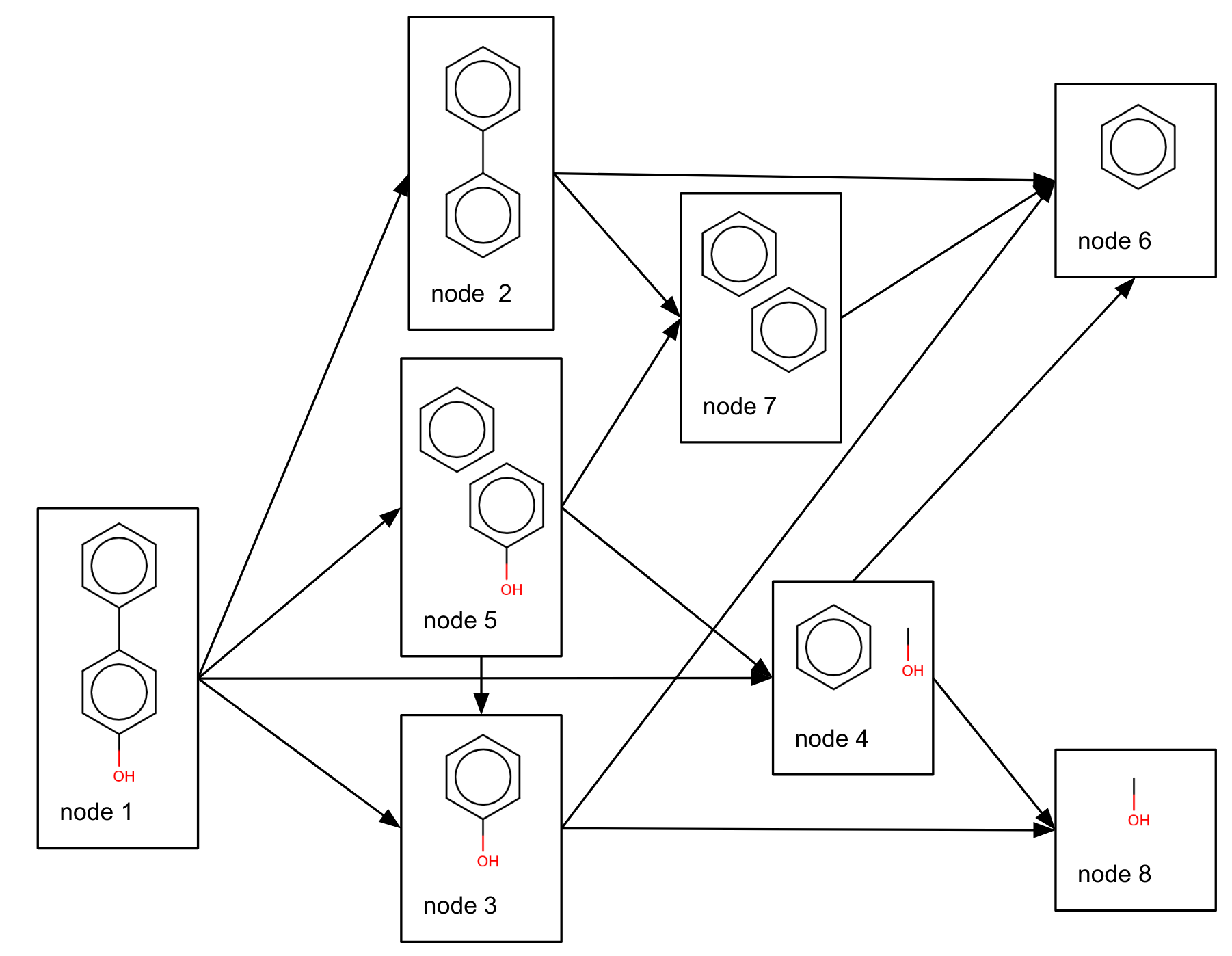
This is repeated for all molecules in your dataset. Of course some molecules will be in common between multiple fragmentation patterns resulting in an extended directed acyclic graph.
The graph can now be traversed to identify related molecules. An edge in the graph is a parent-child relationship and those edges can be traversed in either direction, from a larger parent molecule to a smaller child molecule or vice versa.
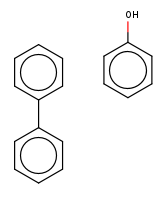
If you consider a single traversal you have simple additions or deletions, such as some chloro additions:
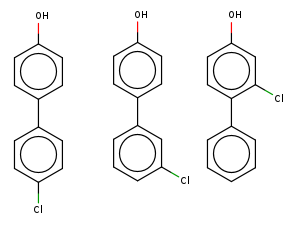
or for instance some deletions:
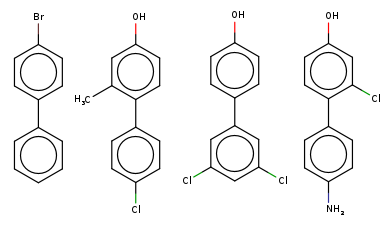
If you now consider two traversals of the graph there are many more possibilities including double additions, double deletions, an addition plus a deletion, a substitution:
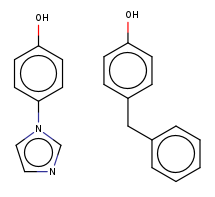
Also, you can have more esoteric changes such as ring substitutions or linker replacements such as these:
To allow these queries to be executed the fragment network data is loaded into a Neo4j graph database that allows these complex graph queries to be executed along with some basic parameters such as the change in the number of heavy atoms or ring atoms. The Fragnet Search application allows a user to specify a query, some parameters and then curate the results, ultimately generating list of molecules that can be purchased.
Details of how we generate these databases will be described in a later post.
The Fragnet Search application uses the REST API that can also be used stand-alone to run queries in a more automated manner, allowing this to be included in an full virtual screening workflow. I’ll describe this in more detail in later posts.